Why AI in Medical Imaging Depends on High-Quality Data Pipelines
In today’s world, AI is not just a tool; it’s becoming essential to modern healthcare. AI helps detect early-stage cancers and predicts cardiac risks before symptoms show up. This technology allows doctors to look beyond the obvious and make quicker, life-saving decisions.
However, every intelligent diagnosis from AI starts long before training a model. It starts deep within medical imaging data. Each CT, MRI, or X-ray scan contains thousands of data points that represent the hidden language of the human body. For the human eye, it’s just an image; for AI, it’s valuable knowledge if the data is clean, organized, and precise.
In healthcare AI, if the input is poor, the output will be poor too, and the consequences involve human lives.
Our team has developed expertise in addressing this challenge: we take thousands of complex medical scans and turn them into reliable, production-ready datasets. This article looks at the DICOM post-processing workflow, the unseen structure that ensures medical AI models learn from accurate information, not noise.
If you’re exploring broader AI orchestration challenges, click here to see how enterprise pipelines scale in real-world systems.
Medical Imaging Data: Beyond 2D Images
When you think of a medical scan, you probably imagine a single X-ray or MRI image like a photograph. That’s not quite how it works in practice.
Medical Scans Capture 3D Data, Not Flat Images
Unlike a camera that captures one 2D frame, medical scanners (CT, MRI, Ultrasound) capture sequences of thin cross-sectional slices stacked together. Imagine slicing an apple from top to bottom; each slice reveals a different layer of the internal structure.
Depth perception: Doctors need to see how organs and tissues are positioned relative to each other across multiple layers
Disease detection: A tumor is not flat; it has depth and shape in three dimensions. To assess its size and seriousness, doctors analyze it across multiple image slices and calculate its volume.
Precise diagnosis: What looks normal in one slice might reveal disease in an adjacent slice
When all these 2D slices are stacked in sequence, they form a complete 3D representation of the patient’s anatomy. Modern AI systems use this 3D structure to understand spatial relationships that 2D analysis would miss.
What is DICOM? The Standard Behind Medical Imaging Data
DICOM metadata processing organizes medical data in a strict hierarchy to prevent confusion and ensure patient safety:
Patient (0010,0010)
└─ Study (0020,000D) – All scans from one hospital visit
└─ Series (0020,000E) – One complete scan sequence
└─ Instances – Individual image slices
└─ Annotations – Radiologist markings (ROIs)
This structure enables:
- Zero patient data mix-ups across the entire imaging workflow
- Preserved clinical meaning as data moves between systems
- Consistent AI training using standardized metadata
- Automatic 3D volume reconstruction in CT and MRI scans
- Global compliance with DICOM PS3 standards
Types of Medical Imaging Modalities and Their Use Cases
Different medical conditions require different scanning technologies:
| Modality | Technology | Use Case |
|---|---|---|
| X-Ray | Electromagnetic radiation | Bones, chest imaging. It is fast and low-cost |
| CT Scan | Computed tomography (multiple angles) | Internal organs, injuries, and tumor detection |
| MRI | Magnetic fields | Brain, spine, soft tissue; excellent for detail without radiation |
| Ultrasound | Sound waves | Real-time imaging, fetal monitoring, and abdominal exams |
| PET Scan | Positron emission | Cancer detection, metabolic activity assessment |
Each modality captures different types of clinical information. An MRI is useless for detecting bone fractures, while an X-ray can’t assess soft tissue damage. The DICOM file should correctly identify which modality was used, and this single field determines how the entire dataset should be processed.
From Annotation to AI: The Medical Image Segmentation Workflow
Transforming raw medical imaging data into AI-ready datasets is a meticulous, multi-step process that ensures accuracy, consistency, and reliability. From cleaning and standardization to segmentation and compliance, each stage plays a critical role in enabling trustworthy and clinically meaningful AI outcomes.
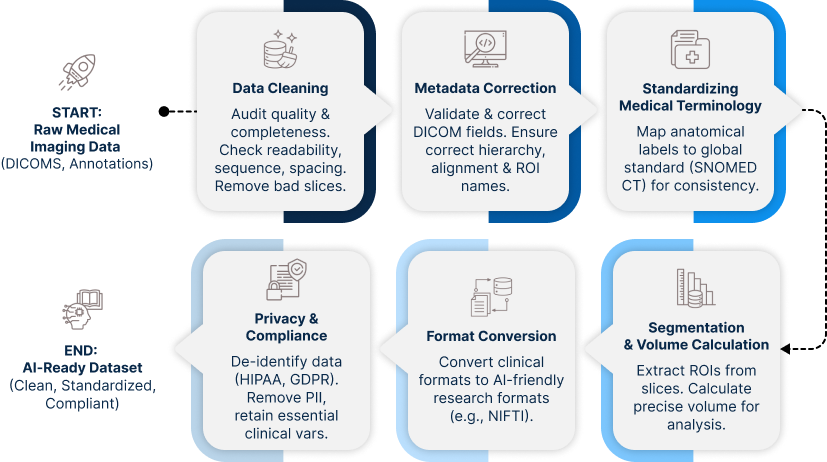
Here’s the journey a medical imaging dataset takes before it’s ready for AI model training:
Step 1: Data Cleaning
Before any analysis happens, the dataset must be audited for quality and completeness to ensure data quality in medical AI.
What we check:
- Are all DICOM files readable? (Corrupted files are discarded)
- Do slices follow the correct anatomical order?
- Are spacing and orientation consistent within each series?
- Is the pixel data within expected intensity ranges?
- Are any slices duplicated or missing?
Why this matters: A single corrupted slice embedded in 500 good slices might not cause an obvious error, but could systematically bias AI model predictions. Finding and removing these problems early prevents downstream disasters.
Step 2: Metadata Correction
Every DICOM file has two main components:
- Pixel Data – the actual scan image slices
- Metadata – stored in the form of Key-Value pairs called DICOM tags
DICOM metadata is complex and a single file contains hundreds of metadata fields such as Study Instance UID, Series Instance UID, Frame of Reference UID, and dozens more. Each one has a specific purpose.
Key metadata fields we validate:
- (0010,0010) PatientName: Patient identifier (anonymized for research)
- (0008,0060) Modality: Scan type (CT/MRI/US/etc) must match actual scan technology
- (0008,103E) SeriesDescription: Human-readable description of what was scanned (e.g., “Chest CT with contrast”)
- (0020,000D) StudyInstanceUID: Links all scans from one visit together
- (0020,000E) SeriesInstanceUID: Groups all slices forming one scan
- (3006,0026) ROIName: Organ or lesion being annotated (e.g., “Liver,” “Kidney Mass”)
- (0020,0052) FrameOfReferenceUID: Ensures all slices stay aligned in anatomical space
Why corrections are essential:
If any of these tags are missing, incorrect, or inconsistent, then the annotations may not match the right scan. Volume and measurement calculations can become inaccurate, and 3D reconstruction may fail due to misaligned slices. So, the AI model might learn incorrect anatomical patterns, and patient follow-up across multiple time points cannot be tracked.
Step 3: Standardizing Medical Terminology
Radiologists around the world use different terms for the same anatomical structures. One might write “Left Kidney Cortex,” another might write “L Kidney Cortical Region.”
To enable consistent AI training across institutions, we standardize these labels using SNOMED CT (Systematized Nomenclature of Medicine Clinical Terms), a global medical terminology standard.
Example:
“Left Kidney Cortex” (radiologist annotation)
↓ (standardized to)
SNOMED CT Code: 181414003
Benefits:
- Cross-institution consistency: Hospitals worldwide train on the same standardized labels
- No ambiguity: Code 181414003 always means the same thing, regardless of language or radiologist preference.
- Better AI interpretation: Models learn from cleanly standardized inputs, not human variation
Step 4: Segmentation and Volume Calculation
For diseases like cancer, precise measurement is critical. Radiologists annotate tumors across multiple slices, but how do we calculate volume?
The process:
- Extract the annotated region, ROI (Region of Interest), from each slice
- Calculate the area of that region in each slice
- Multiply by slice thickness and pixel spacing
- Sum across all slices
Formula:
Volume = Σ (Area of ROI per slice × Slice Thickness × Pixel Spacing)
This sounds simple, but precision matters the most. A 5% error in volume calculation could change treatment decisions.
Step 5: Format Conversion
Medical imaging uses multiple file formats for different purposes:
- DICOM (.dcm): Standard clinical format with full metadata
- RTSTRUCT (.dcm): Radiotherapy structure sets annotations stored separately from image data
- DICOM SEG (.dcm): Segmentation objects in modern DICOM format
- NIfTI (.nii.gz): Medical research format, compact and AI-friendly
AI training pipelines often need data in NIfTI or segmentation mask format. Converting between formats while preserving accuracy is a specialized skill, and one wrong step corrupts the data.
Step 6: Privacy and Compliance
Healthcare data is legally protected under HIPAA (USA), GDPR (Europe), and NDHM (India). The dataset must be de-identified before research use by removing any personally identifiable information.
What gets removed:
- Patient name and ID
- Date of birth
- Institution name
- Any text that could identify the patient
What stays (essential for AI):
- Age or age range
- Gender
- Scan type and modality
- Anatomical location
- Clinical findings
Balancing de-identification with data usefulness is the challenge. Remove too much, and the dataset becomes useless. Keep too much, and you’ve violated privacy regulations.
See Also, Medical Images Annotation
The Challenges in the Workflow
The DICOM post-processing is not complex due to any single factor; it’s complex because many factors must align perfectly simultaneously.
The Scale Problem
A typical clinical study containing 500 slices of CT means:
- 500+ metadata verification steps
- Over 500 slice alignment checks
- Multiple volume calculations (one per annotated organ/lesion)
- Everyone must be precise
Scale up to thousands of studies, which is needed for robust AI training, and the challenge becomes managing consistency at scale.
The Cascade Effect
Errors do not happen in isolation. One metadata error might corrupt:
- 3D reconstruction (slices won’t align)
- Volume calculations (wrong anatomical space)
- Training the AI model (wrong signal)
- Clinical interpretation (misdiagnosis support)
It is necessary to catch errors at the source to prevent cascade failures from going downstream.
The Format Fragmentation Problem
Different institutions use different DICOM conformance levels. Some include all recommended metadata; others skip optional fields. Conversion between formats (RTSTRUCT → DICOM SEG → NIfTI) compounds the challenge, with each conversion being a potential failure point.
Balancing Automation and Human Expertise
Some validation steps are automatable (checking for corrupted files, verifying UID uniqueness). Other steps require a radiologist’s expertise to confirm that an annotation actually represents what it claims. Building pipelines that combine automated checks with expert review and without creating bottlenecks is a design challenge.
Our Approach: Precision at Scale
Our DICOM post-processing workflow is built on three principles:
1. Automation for Consistency
We programmatically validate metadata, check spatial relationships, and convert formats using specialized DICOM processing libraries, pydicom, DCMQI, and SimpleITK. Automation catches issues that manual review would miss.
2. Expert Validation for Nuance
Automated systems can flag suspicious data, but human radiologists make final determinations. We combine algorithmic checking with clinical expertise: the best of both worlds.
3. Compliance by Design
It means no afterthoughts regarding privacy, DICOM standards, or audit trails; these are embedded into the pipeline. De-identification, HIPAA/GDPR compliance, and compliance verification at each step happen automatically.
Result: datasets that are clean, standardized, compliant, and ready for trustworthy AI model training.
Why Investment in Post-Processing Pays Off
It might seem like overkill to spend this much effort cleaning data when you could just throw raw scans into an AI training pipeline. But consider the alternative:
Scenario 1: Rushing Model Development
When raw, uncleaned data goes straight into model training, the AI learns from corrupted or inconsistent inputs. It might look good during testing, but fail in real-world use, causing hospitals to lose trust. This can risk patient safety, trigger regulatory scrutiny, and ultimately lead to project failure.
Scenario 2: Investing in Data Quality
When the data is properly cleaned and validated, the AI model learns from reliable information. It performs consistently in both testing and production, leading hospitals to adopt it with confidence. The result is better clinical outcomes, regulatory compliance, and a system that’s built to last.
The Lesson
Poor data quality doesn’t just cause system errors, and it erodes trust and can put patients at risk.
Conclusion: The Foundation of Medical AI
The advancement of medical AI depends less on algorithms and more on data integrity. While cutting-edge models attract attention, the critical work of data cleaning, metadata correction, standards compliance, and volume validation determines whether AI systems can be deployed safely in clinical settings. Without rigorous data preparation, even the most advanced algorithms remain experimental rather than practical tools for healthcare delivery.
In today’s healthcare technology, competitive advantage stems from data quality, not just model complexity. Organizations that establish robust processes for verifying DICOM tags, aligning imaging data, validating calculations, and ensuring patient data protection create the foundation for AI systems that clinicians can trust. This precision-focused approach transforms AI from a promising concept into a reliable clinical asset.
Our proven methodology centers on this fundamental principle: medical AI must be built on verified, standardized, and meticulously maintained data. By prioritizing data integrity at every stage from initial collection through processing and deployment, we enable AI systems that meet the rigorous standards healthcare requires.
In an industry where accuracy can mean the difference between effective treatment and patient harm, data quality is not merely a technical requirement but an ethical imperative.
Author’s Note: This article was supported by AI-based research and writing, with Claude 4.5 assisting in the creation of text and images.
What is DICOM in medical imaging?
DICOM (Digital Imaging and Communications in Medicine) is the standard format used to store and transmit medical imaging data along with metadata, ensuring consistency across healthcare systems.
Why is data quality important in medical imaging AI?
Poor-quality data leads to inaccurate AI predictions, which can impact diagnosis and patient safety. Clean, validated datasets ensure reliable model performance.
What is DICOM post-processing?
DICOM post-processing involves cleaning, validating, standardizing, and converting imaging data to make it usable for AI training and clinical applications.
How does AI use 3D medical imaging data?
AI models analyze stacked 2D slices as 3D volumes to understand spatial relationships, detect abnormalities, and measure disease progression.
What is medical image segmentation?
Segmentation is the process of identifying and isolating regions of interest (like tumors) in imaging data to enable precise measurement and analysis.
How is patient privacy maintained in medical AI datasets?
Sensitive data is removed through de-identification processes while retaining essential clinical information, ensuring compliance with regulations like HIPAA and GDPR.

