Introduction: Rethinking Data Extraction
Clinical literature is the lifeblood of pharmaceutical research, but also one of its biggest bottlenecks. That is to say, extracting structured insights from trial publications can require weeks of manual review, as human experts search through dense narratives, tables, and figures.
Working in partnership with top-20 pharma manufacturers, we set out to reimagine this process. In this regard, our platform is built to apply AI not just as a helper, but as a transformational layer for parsing and structuring clinical intelligence.
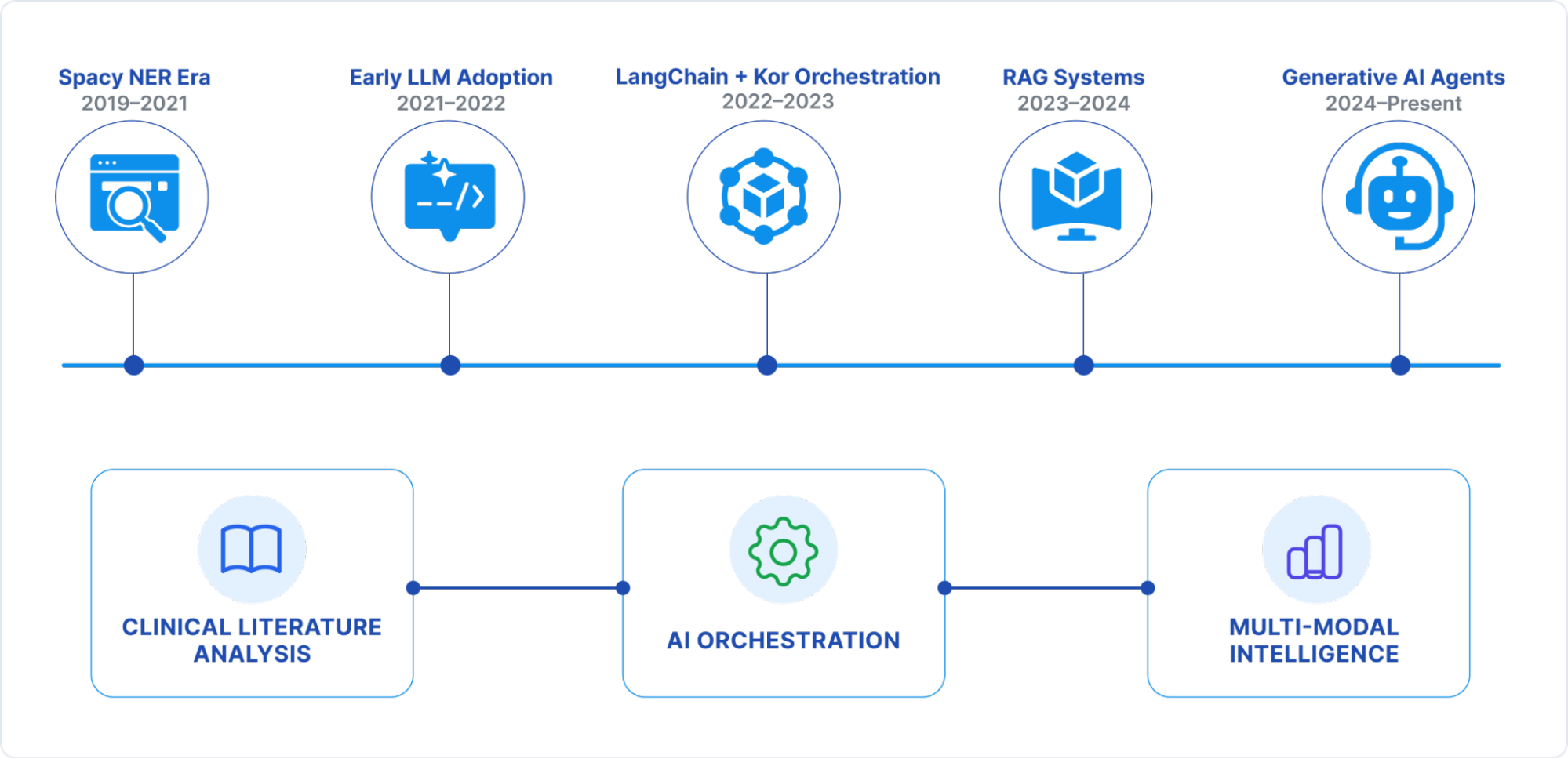
Our journey over the past five years mirrors the broader evolution of NLP from traditional rule-based NER to LLM-powered agents with multi-modal capabilities. In this blog, we share that evolution: the challenges, architectural shifts, measurable gains, and lessons learned along the way.
Stage 1: The Spacy NER Era (2019–2021)
In Stage 1, the extraction pipelines leaned on custom Spacy-based NER models, trained to recognize clinical trial entities such as drug names, study endpoints, and patient cohorts.
Specifically, the Architecture included:
- Statistical entity recognition models
- Rule-based post-processing and validation
- Entity linking against medical vocabularies like MeSH
However, several challenges emerged:
- Annotation overhead, that is, months of expert effort to build domain datasets
- GPU-heavy infrastructure for real-time inference
- Constant retraining cycles for new domains
As a result, performance was as follows:
- Accuracy: 65–75% on core entities
- Throughput: 2–3 docs/min per GPU
While limited in scope, this phase laid the groundwork for structured data pipelines and showed that automation could meaningfully augment human reviewers.
Stage 2: Early LLM Adoption (2021–2022)
The arrival of GPT marked an inflection point. Consequently, by leveraging APIs for few-shot prompt-driven extraction, we bypassed rigid training pipelines.
As a result, several things changed:
- No more week-long annotation cycles, contextual reasoning of LLMs filled the gap
- JSON-structured extraction via prompt engineering
- Generalization across clinical subdomains
This led to a measurable impact:
- Accuracy jumped to ~80%
- Additionally, a 60% reduction in manual annotation effort
- Deployment cycles compressed from months to weeks
Overall, this was our first taste of LLMs as adaptable engines rather than narrow models.
Stage 3: Structured Orchestration with LangChain + Kor (2022–2023)
Direct LLM calls worked, but at production scale, orchestration was critical. Therefore, we introduced LangChain for workflow management, and later Kor for schema enforcement.
Engineering Innovations:
- Reusable prompt templates and chains
- Built-in error handling and retries
- Moreover, Kor for strict schema validation using Pydantic
Impact:
- Consistency in data structure jumped to 85%
- Throughput up by 40%
- Error rates cut by 30%
For the first time, we achieved production-grade reliability rather than one-off model experiments.
Stage 4: Retrieval-Augmented Generation (2023–2024)
Clinical literature often hides meaning in contextual fragments across disparate sources. To solve this, we embedded corpora into vector databases, enabling RAG-driven context injection into model prompts.
Architecture Highlights:
- Semantic search over domain embeddings
- Multi-document reasoning for trial reports
- Reduced model hallucination in dense medical contexts
Results:
- Accuracy surged past 90% for complex relationships
- Multi-page trial parsing became coherent
- Furthermore, terminology disambiguation (abbreviations, synonyms) dramatically improved
In essence, RAG lets models “think” with knowledge in hand, not guess.
Stage 5: Generative AI Agents (2024–Present)
Today, our application employs multi-agent systems that are specialized autonomous units for different data modalities and clinical domains.
Features:
- Task-oriented agents (treatment arm, safety data, biomarkers)
- Self-correction and validation agents in the loop
- Multi-modal inputs: text + tables + figures
What’s Possible Now:
- Extracting granular dosing regimens and patient stratification
- Parsing clinical charts, Kaplan–Meier curves, and molecular pathways
- Temporal + causal reasoning across trial timelines
Performance:
- Accuracy: 90%+
- Processing Speed: 15–20 docs/min
- Meanwhile, the annotation needs to be cut by 90%
- Processing costs down by 60% (CPU-based serverless infra)
Currently, the platform acts as a domain-aware research assistant, not just an extraction engine.
Lessons Learned: Building AI for Clinical Research
- Evolve with the ecosystem: Rapid LLM advances forced constant reassessment. Betting on modular, API-first architecture lets us adapt quickly.
- Data quality is paramount: Automated schema validation + human-in-loop review were essential to win trust.
- Design for scale, not pilots: From GPUs to cloud-native serverless infra, scalability had to be baked in.
- Multi-modality is non-negotiable: Clinical data resides in tables and figures, not just text.
The Roadmap: Beyond Text Extraction
Looking ahead, the future lies in real-time, multi-modal clinical intelligence pipelines:
- Next-gen biomedical LLMs optimized for trial data
- Video and audio parsing from medical presentations
- Real-time monitoring of ongoing clinical trials
- Seamless integration with regulatory and compliance frameworks
In conclusion, the roadmap is clear: from extraction to interpretation, from static reports to dynamic clinical intelligence.
Author’s Note: This article was supported by AI-based research and writing, with Claude 4.4 assisting in the creation of text and images.

